- EGFR, a receptor tyrosine kinase, plays an important role in cell proliferation by activating downstream signaling pathways, including RAS4
- Inactivation of the mutant RAS signaling pathway leads to reactivation of EGFR and subsequent lateral stimulation of wild type RAS pathways1-3
KRASG12C, an altered KRAS protein due to a RAS mutation, drives cancer cell growth and survival5
See below for a deeper look into the role of KRAS G12C in cancer6
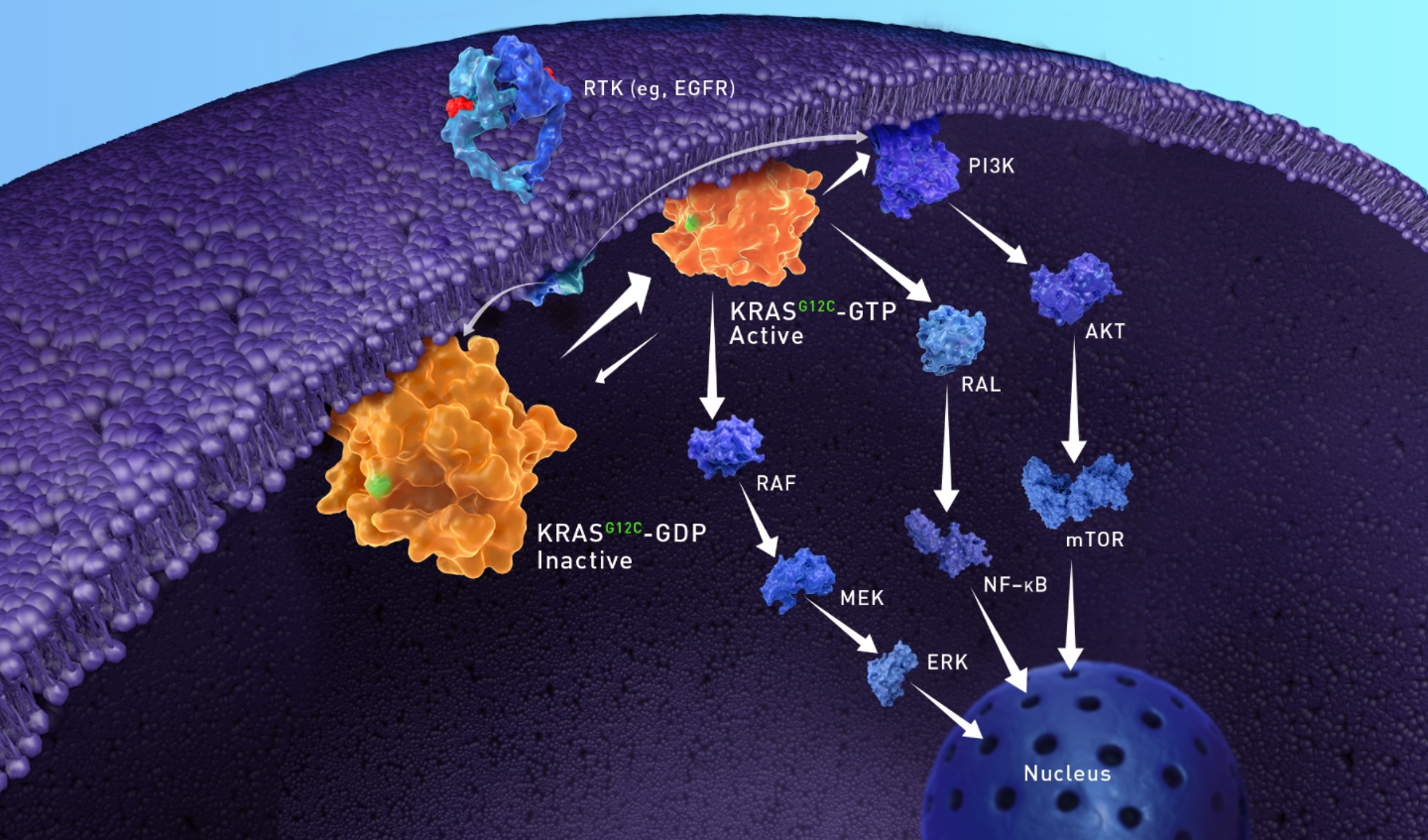

KRAS G12C oncogenic signaling
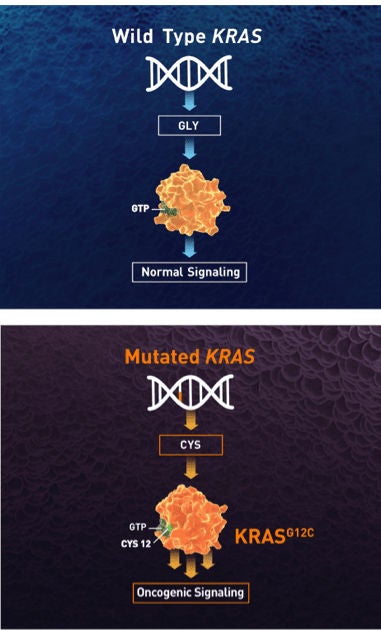
- In normal cells, KRAS cycles between an active and inactive state to regulate downstream signaling pathways, leading to cellular proliferation, differentiation, and survival7,8
- In tumor cells, the active GTP-bound form of KRASG12C is favored, resulting in oncogenic signaling and tumorigenesis5
- KRASG12C is a glycine-to-cysteine amino acid substitution driven by a point mutation in the KRAS coding sequence9
Comprehensive RAS testing includes KRAS and detects mutations such as KRAS G12C.
Having these results accessible can help inform clinical decisions10
AKT, protein kinase B; CYS, cysteine; EGFR, epidermal growth factor receptor; ERK, extracellular signal-regulated kinase; GDP, guanosine diphosphate; GLY, glycine; GTP, guanosine triphosphate; KRAS, Kirsten rat sarcoma; MEK, mitogen-activated protein kinase kinase; mTOR, mammalian target of rapamycin; NF-κB, nuclear factor kappa-light-chain-enhancer of activated B cells; PI3K, phosphoinositide 3-kinase; RAF, rapidly accelerated fibrosarcoma; RAL, Ras-like; RAS, rat sarcoma; RTK, receptor tyrosine kinase.
References: 1. Henry JT, et al. JCO Precis Oncol. 2021;5:613-621. 2. Amodio V, et al. Cancer Discov. 2020;10(8):1129–1139. 3. Feng J, et al. Oncogene. 2023;42(20):1620-1633. 4. Uribe ML, et al. Cancers (Basel). 2021;13(11):2748. 5. Ryan MB, et al. Nat Rev Clin Oncol. 2018;15(11):709-720. 6. Neel NF, et al. Genes Cancer. 2011;2(3):275-287. 7. Merz V, et al. Front Oncol. 2021;11:638360. 8. Ferrer I, et al. Lung Cancer. 2018;124:53-64. 9. Ihle NT, et al. J Natl Cancer Inst. 2012;104(3):228-239. 10. Roche Diagnostics. https://diagnostics.roche.com/us/en/products/params/cobas-kras-mutation-test.html. Accessed June 9, 2023.